Comparative Genomics of Human Pathogens
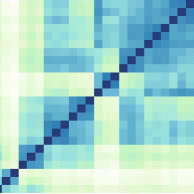
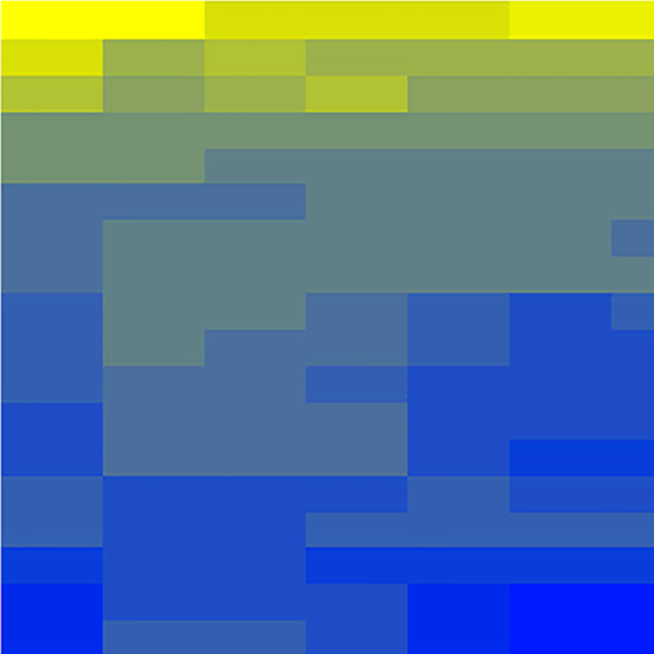
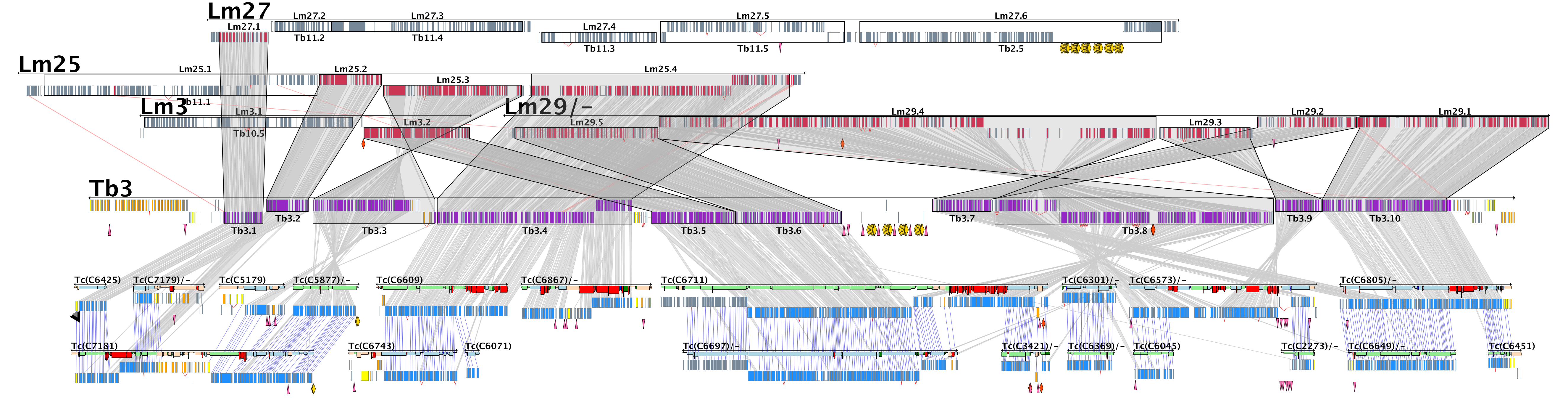
A wide range of pathologies is found within the lineages of trypanosomatids and there is a wide range of evolutionary and pathological space yet to be explored through additional comparative sequencing and analyses. A special area of interest within our group is the comparative genomics of pathogens with an emphasis on the study of the evolution of genome architecture and content, as well as the identification of genes that are phylogenetically restricted. Our recent investigations have focused on the three trypanosomatid parasites T. brucei, T. cruzi, and L. major. The three pathogens are related but have distinctive life cycles, different insect vectors, and, perhaps most importantly, different disease characteristics.
Our current comparative efforts encompass a variety of pathogenic and non-pathogenic trypanosomatid strains for which sequencing projects are underway. Comparative analyses of these genomes will not only reveal great insights into the evolution of trypanosomatids, but will allow us to identify subsets of genes specifically linked to virulence and survival mechanisms. We are poised to make contributions to the effort aimed at answering several outstanding questions in the pathogenesis of trypanosomatids: What is/are the genotypes associated with the ability of different strains or isolates to cause widely varied clinical manifestations? Are certain metabolic, regulatory, or genetic networks required for or associated with disease? What are the genetic bases of the phenotypic (cell cycle, host range, vector selection, pathogenic and clinical manifestations, etc.) characteristics of the major groups of pathogenic trypanosomatids?